Multi-locus GWAS of Ethiopian sorghum landraces revealed extensive genetic variation and key candidate genes underlying root architectural traits, providing valuable targets for breeding drought-resilient, water-efficient sorghum cultivars.
Keywords: BiomaRt, LOD scores, ML-GWAS, Nodal root angle, QTNs, RSA, SNP
Ethiopia, a primary center of origin for sorghum, harbors extensive genetic diversity that underpins the crop’s resilience to abiotic stresses such as drought. This diversity offers an invaluable foundation for identifying genetic variants linked to drought adaptation, particularly root system architectural (RSA) traits critical for water acquisition. Through advances in high-throughput genotyping and multi-locus genome-wide association studies (ML-GWAS), researchers can now dissect the complex polygenic basis of traits like nodal root angle, root length, and root number with improved precision. In Ethiopian sorghum landraces, researchers from Addis Ababa University, Melkassa Agricultural Research Center, Jimma University and collaborating institutions used ML-GWAS to reveal rapid linkage disequilibrium decay, indicating a high level of genetic diversity and enabling the detection of 73 quantitative trait nucleotides (QTNs) distributed across all sorghum chromosomes. Notably, QTNs associated with root angle explained up to 15% of phenotypic variation, highlighting both the genetic complexity of this trait and its potential for marker-assisted selection (MAS).
Functional annotation and enrichment analyses revealed that candidate genes regulating RSA traits are involved in key biological processes, including transcriptional regulation, DNA repair, and hormonal signaling. Genes such as protein phosphatase 2C (PP2C) and cytochrome P450 were linked to drought-responsive pathways, while others, including E3 ubiquitin-protein ligase BRE1 and histone lysine demethylases, pointed to epigenetic mechanisms modulating stress responses. Correlations among RSA traits, such as between nodal root angle, root length, and plant height, underscore their coordinated control and pleiotropic genetic basis. Collectively, these findings illuminate the genetic architecture underlying drought tolerance in Ethiopian sorghum landraces and establish a genomic framework for developing climate-resilient, water-efficient sorghum cultivars through integrative breeding strategies.
SorghumBase Examples:
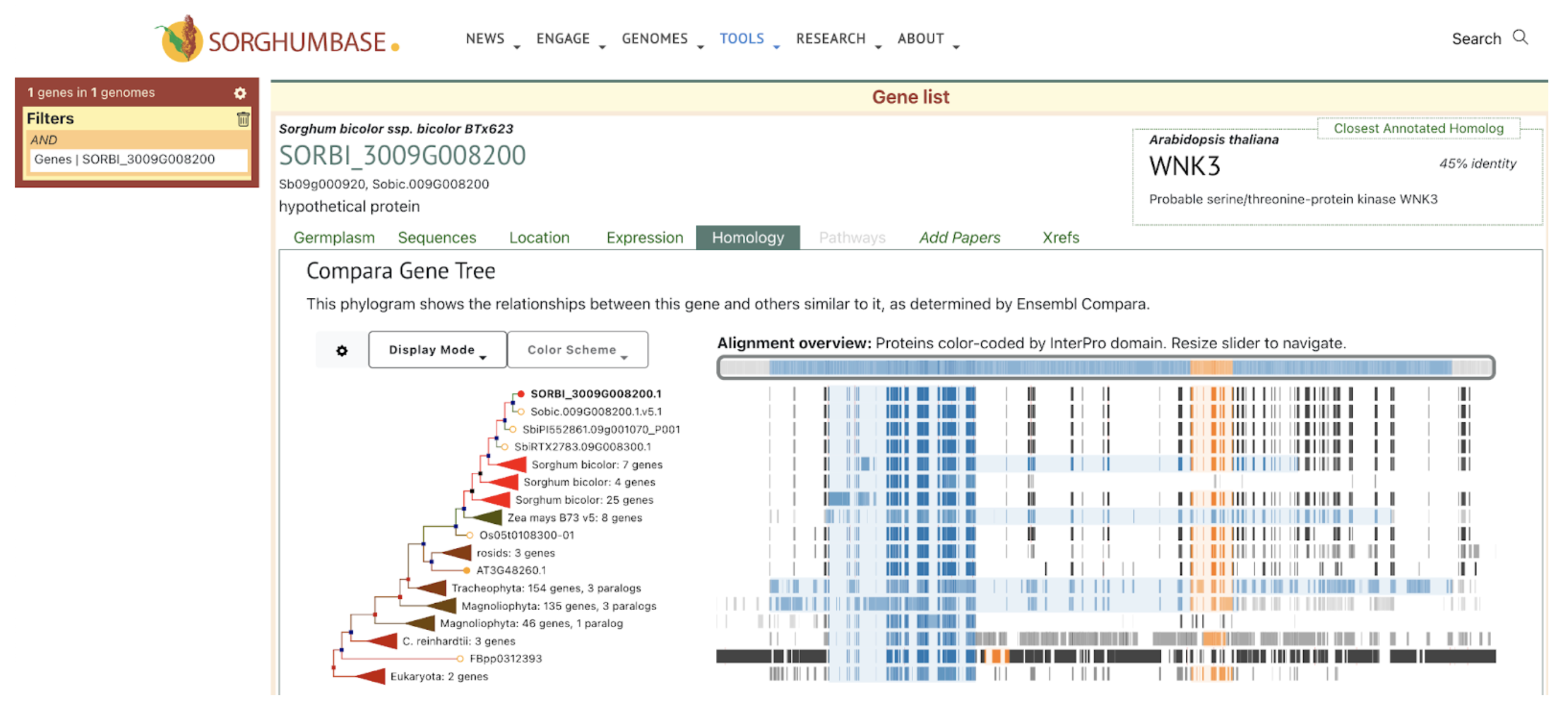
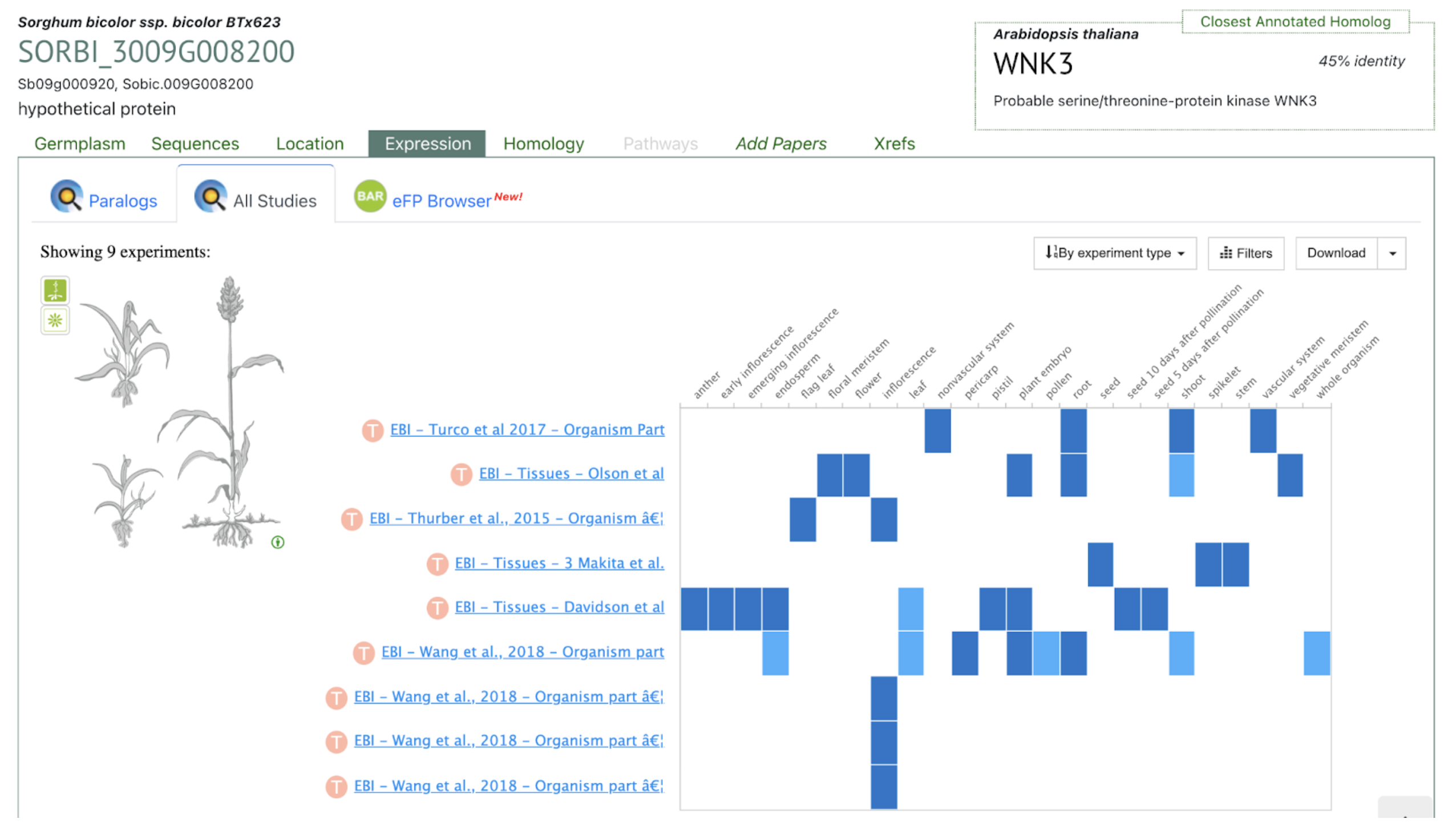
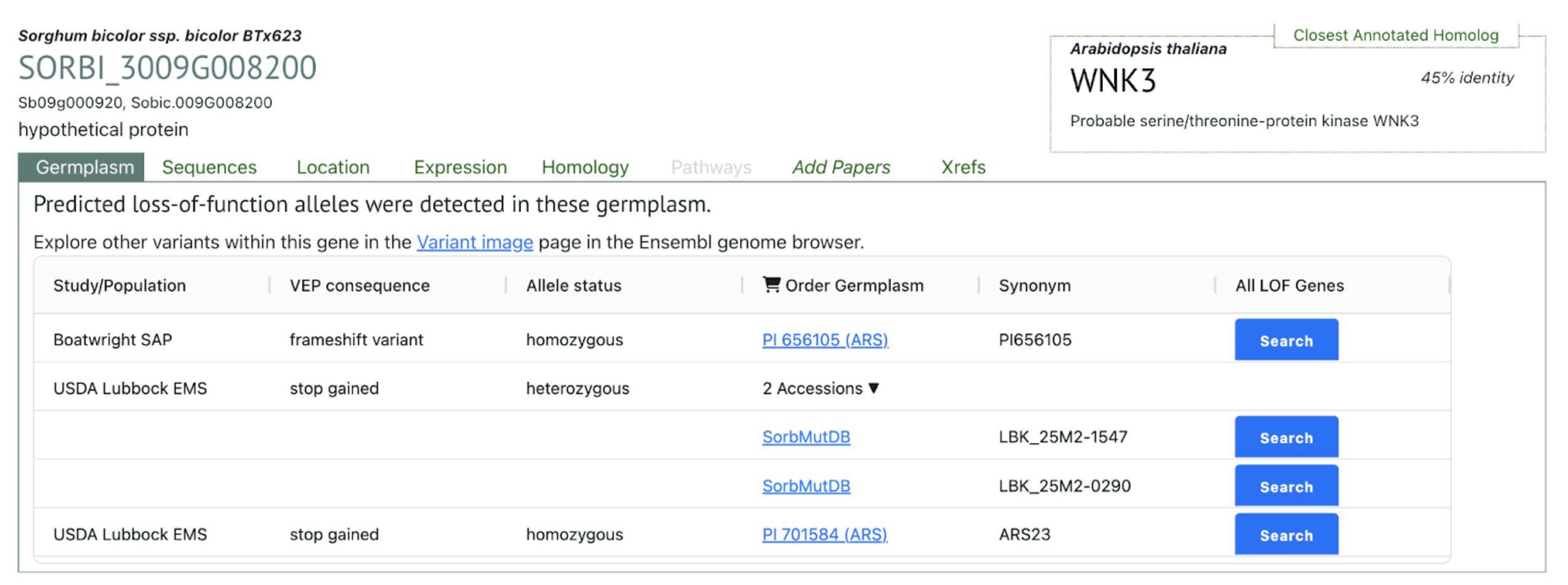
Mitiku AD, Feyissa T, Woldetensaye AT, Chikssa HN, Menamo TM, Abebe TM, Bante K. Multi-locus genome-wide association studies for root system architectural traits in Ethiopian sorghum (Sorghum bicolor L.) landraces. BMC Plant Biol. 2025 Sep 2;25(1):1180. PMID: 40890595. doi: 10.1186/s12870-025-07271-6. Read more