Sorghum genotypes exhibit adaptations to combined drought and salinity stress, through enhanced antioxidative defense, osmotic adjustment, and stress-responsive gene expression.
Keywords: Sorghum bicolor, abiotic resilience, genetic variation, oxidative stress mitigation, sustainable agriculture, targeted breeding
Researchers from King Abdulaziz University and Federal University Dutsinma investigated the integrated stress responses of sorghum to combined drought and salinity stress, revealing complex interactions among physiological, biochemical, and molecular mechanisms. Antioxidant enzyme activities, including superoxide dismutase (SOD), peroxidase (POD) and catalase (CAT), varied among genotypes, highlighting genotype-specific defense strategies. Notably, CRS-01 exhibited lower SOD but higher CAT activity, suggesting differential oxidative stress regulation. Osmolyte accumulation, such as proline and glycine betaine, played a critical role in osmotic adjustment. Sorghum genotypes CRS-01 and Samsorg-42 demonstrated superior accumulation of osmolytes under stress conditions. Physiological parameters, including chlorophyll content, Na/K ratio, relative water content (RWC), and water potential, provided insights into stress tolerance, with CRS-01 maintaining favorable levels under severe drought and salinity stress. The decline in photosynthetic rate (Pn) and stomatal conductance (Gs) with increasing stress severity further underscored the trade-off between water conservation and carbon assimilation. However, Samsorg-17 exhibited consistently higher Pn and Gs, indicating its inherent resilience to abiotic stress.
At the molecular level, gene expression analysis highlighted the upregulation of key stress-related genes, including SbSOD1, SbAPX2, SbCAT3, SbHKT1; 4, SbDREB2A, SbDHN3, and SbPRP1, with the highest expression under combined stress conditions. Samsorg-17 displayed the strongest upregulation of these genes, correlating with its superior physiological and biochemical performance. Correlation and principal component analyses (PCA) revealed strong interdependencies among antioxidative defense mechanisms, osmotic regulation, and stress tolerance traits, emphasizing their collective role in mitigating oxidative stress and maintaining cellular homeostasis. Overall, these findings provide valuable insights into the adaptive strategies of sorghum under abiotic stress, guiding breeding efforts to develop more resilient cultivars for sustainable agriculture in arid and saline environments.
SorghumBase examples:
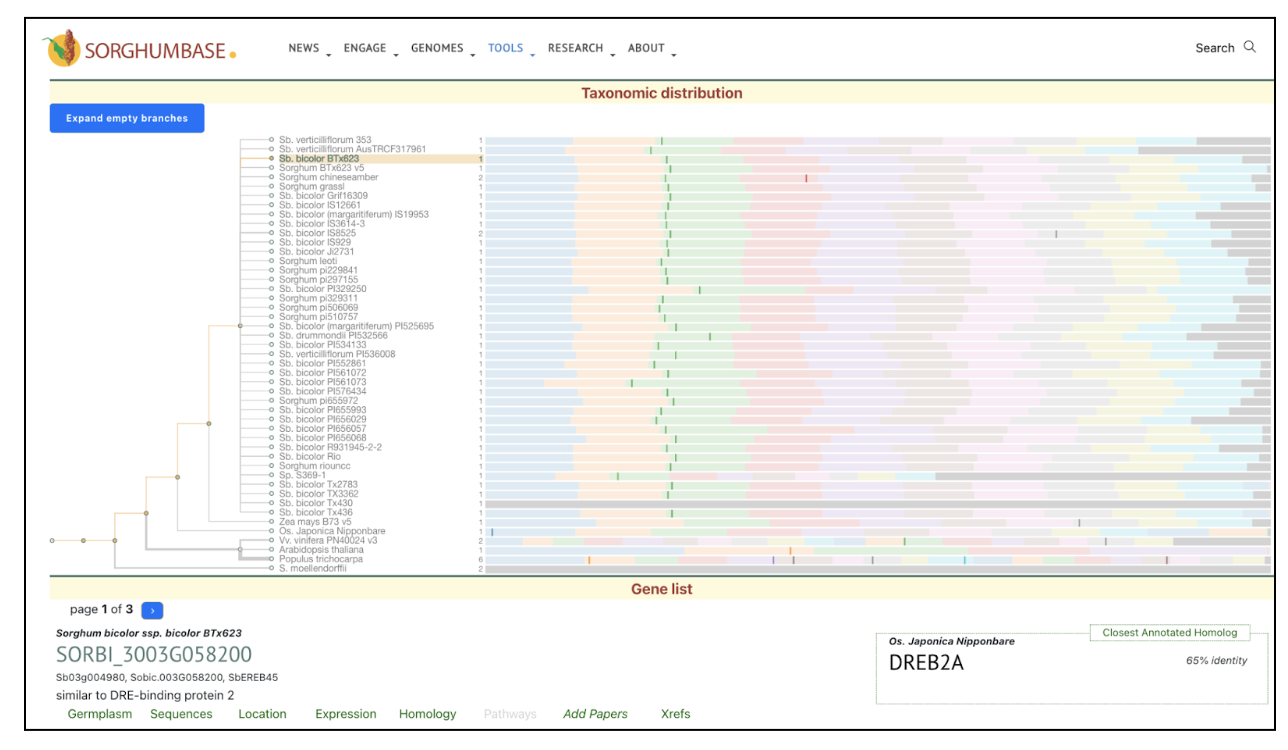
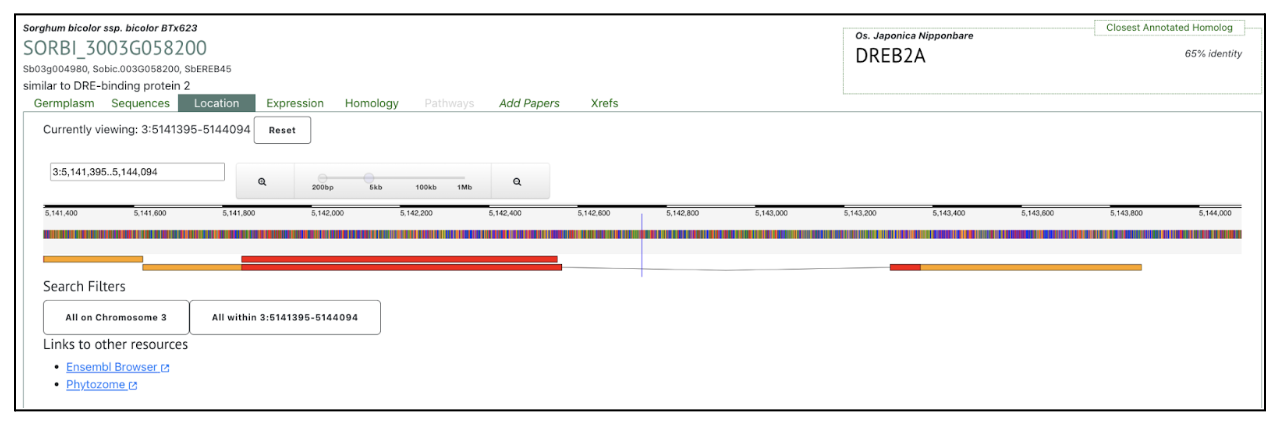
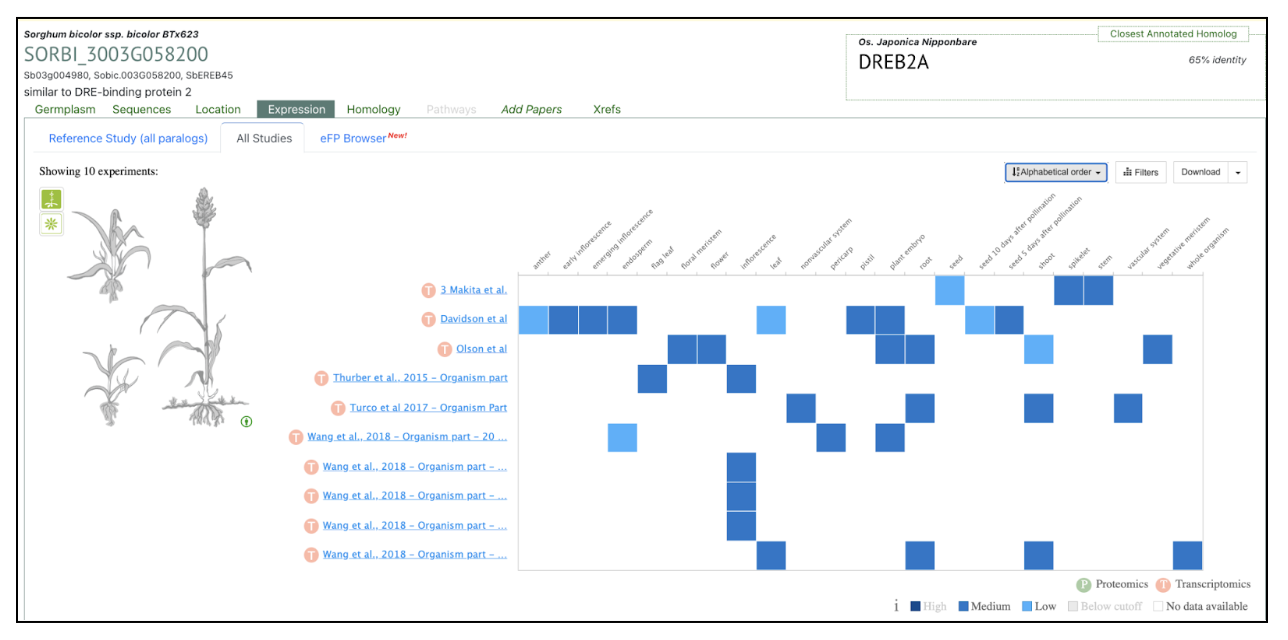
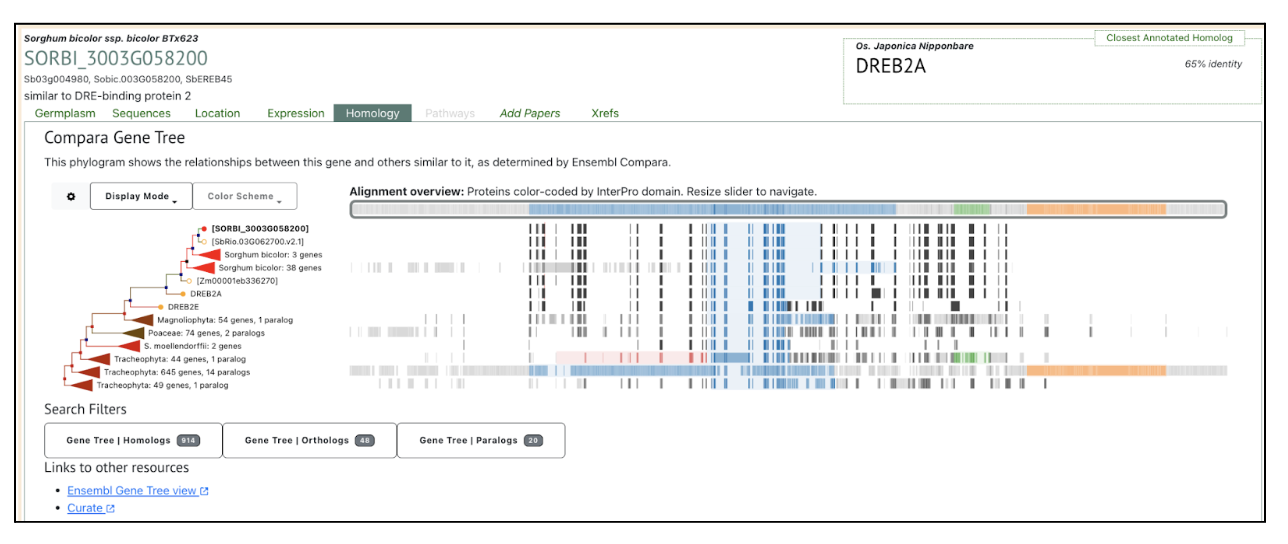
This study demonstrates that SbDREB2A transcription factor is involved in enhancing stress tolerance by regulating gene expression in response to drought and salinity. The DREB2A transcription factor was used as a keyword to search SorghumBase and the taxonomic distribution of the rice DREB2A gene provided its sorghum ortholog “SORBI_3003G058200”. This gene SORBI_3003G058200 was further used to explore its homology in other sorghum accessions and its expression across studies.

Reference:
Alzahrani Y, Abdulbaki AS, Alsamadany H. Genotypic variability in stress responses of Sorghum bicolor under drought and salinity conditions. Front Genet. 2025 Jan 8;15:1502900. PMID: 39845188. doi: 10.3389/fgene.2024.1502900. Read more
Related Project Websites:
- Dr. Alsamadany’s page at King Abdulaziz University: https://halsamadani.kau.edu.sa/CVEn.aspx?Site_ID=0005370&Lng=EN