Conserved non-coding sequences (CNS) play a crucial role in maize gene regulation by interacting with chromatin accessibility and epigenetic modifications, offering potential targets for crop improvement through molecular breeding.
Keywords: Conserved non-coding sequences, Maize, Phenotypic variations, Regulatory Elements
Conserved non-coding sequences (CNS) play a crucial role in gene regulation despite not encoding proteins. This study aimed to identify CNS in maize (Zea mays B73) by comparing its genome with related species, including Zea mays spp. mexicana, sorghum, foxtail millet, and adlay. The analysis conducted by scientists from Sichuan Agricultural University, Sichuan Tianfu New Area Rural Revitalization Research Institute and Chengdu Normal University revealed that 99.09% of CNS are located in intergenic and promoter regions, suggesting their involvement in transcriptional regulation. CNS near transcription start sites (TSS) may function as enhancers or promoters, while those further away may act as long-range regulatory elements. The study also found that CNS overlap with open chromatin regions, transcription factor binding sites, and epigenetic modifications such as H3K9ac, which is associated with transcriptional activation. Additionally, variations in CNS were significantly linked to maize root traits and agronomic characteristics, reinforcing their regulatory potential in plant development and adaptation.
The study further highlights the functional importance of CNS in gene expression and phenotypic variation. Previous research in plants has demonstrated that CNS can serve as enhancers, such as those regulating the KNOX1 gene family in Arabidopsis. In maize, CNS were identified within the flowering-time regulatory region of Vgt1 and upstream of ZmCCT10, both enriched with enhancer-associated histone modifications. These findings suggest CNS could be leveraged in crop molecular breeding to fine-tune gene expression, improving traits like drought tolerance and yield stability. By using technologies such as CRISPR/Cas9, breeders can modify CNS without disrupting protein-coding regions, enabling precise trait enhancements while maintaining genetic stability.
SorghumBase examples:
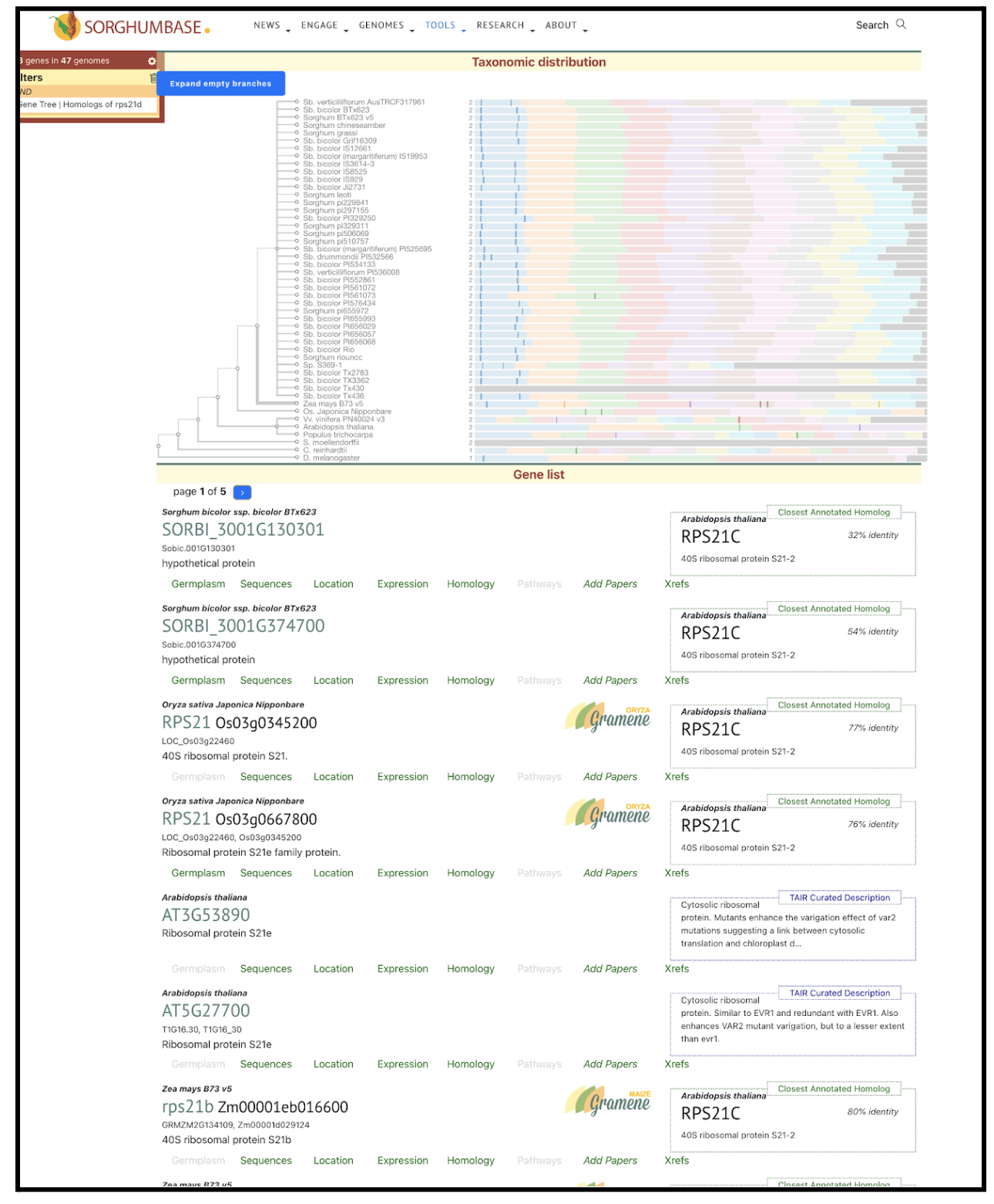
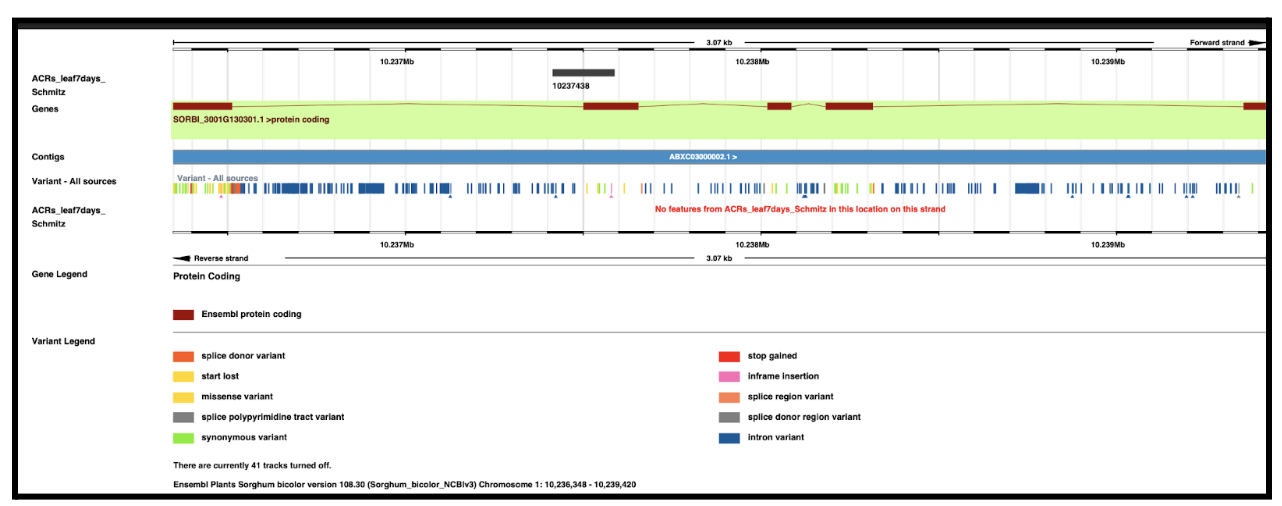
The study identified Zm00001d012886, a maize gene encoding a 40S ribosomal protein, as a key target of conserved non-coding sequences (CNS) that regulate its expression. A total of 41 CNS were found within 5 Kb of the gene, with two SNPs (chr5.S_1568608 and chr5.S_1568631) significantly affecting its expression. These CNS variations were linked to important agronomic traits, including ear leaf length, ear height, and ear length, as well as root traits under drought stress. Gene expression analysis revealed that Zm00001d012886 was more active under drought conditions, suggesting a role in stress adaptation. Since sorghum shares evolutionary similarities with maize, a homologous gene in sorghum may also be CNS-regulated, presenting opportunities for crop improvement in drought-prone environments.

Reference:
Luo Y, Zhai H, Zhong X, Yang B, Xu Y, Liu T, Wang Q, Zhou Y, Mao Y, Liu Y, Tang Q, Lu Y, Wang Y, Xu J. Characterization and functional analysis of conserved non-coding sequences among poaceae: insights into gene regulation and phenotypic variation in maize. BMC Genomics. 2025 Jan 20;26(1):46. PMID: 39833673. doi: 10.1186/s12864-025-11221-9. Read more