Striga resistance in sorghum is achieved through molecular marker-assisted breeding targeting mutations at the LGS1 gene. These lgs1 mutants exude fewer Striga-stimulatory strigolactones, providing a sustainable solution for combating this parasitic weed.
Striga, a parasitic weed, causes substantial economic losses in sub-Saharan African agriculture, particularly in smallholder farms where inputs like improved cultivars, irrigation, and fertilizers are limited. Sustainable control strategies focus on host plant resistance, particularly in sorghum, combined with water conservation and soil nutrition practices. Low-stimulant sorghum lines, such as SRN39, have been crucial for developing resistance. The agar gel assay was an early innovation that enabled breeders to evaluate sorghum root exudates for their ability to stimulate Striga seed germination, uncovering genetic resistance linked to low stimulant activity. These traits, controlled by recessive alleles at the LGS1 locus, were later tracked using molecular markers, significantly enhancing breeding programs. In the present study, researchers from Purdue University developed a robust LGS1 marker to identify lgs1 alleles, which disable a sulfotransferase enzyme critical for producing Striga-stimulatory strigolactones. This marker reliably detects all known lgs1 variants, ensuring precise selection in breeding programs, even in environments unsuitable for direct Striga testing.
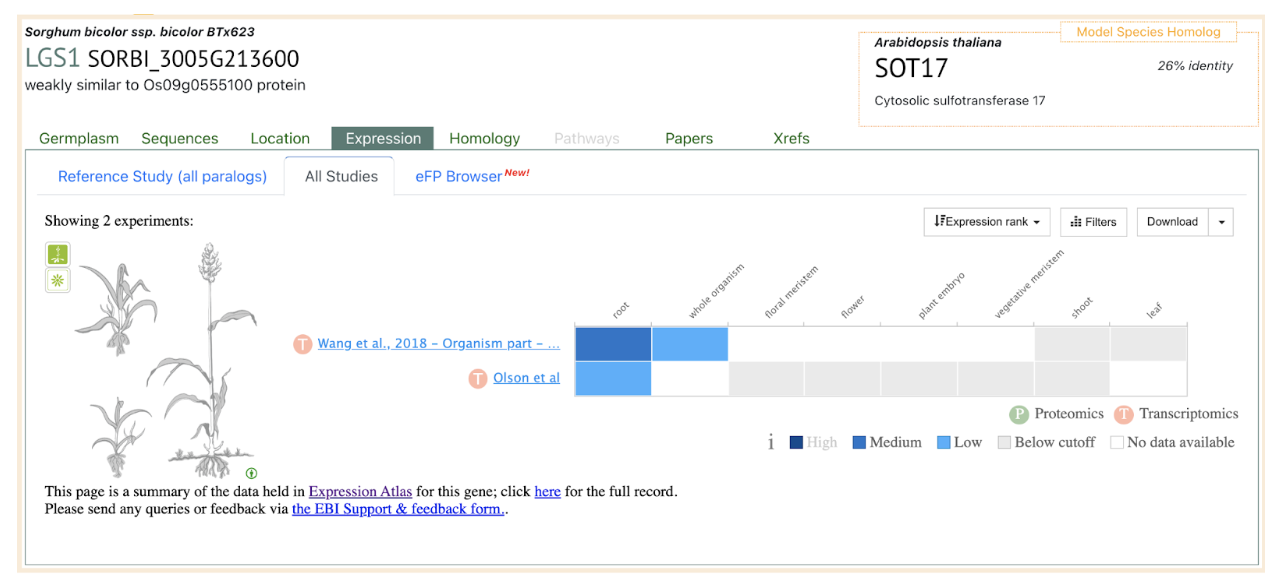
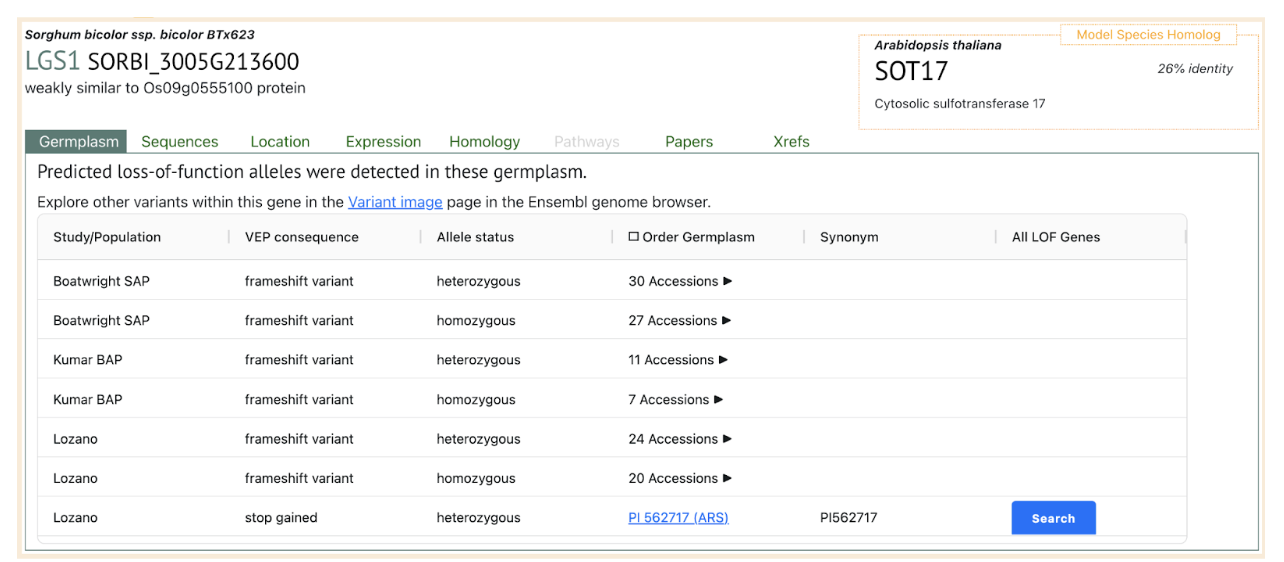
Sequencing revealed significant variation in lgs1 alleles among sorghum lines, with some lines carrying large deletions or smaller loss-of-function mutations at the LGS1 locus. These genetic changes alter strigolactone profiles, reducing Striga germination and improving resistance. Testing of 406 sorghum lines using the new marker identified 23 with lgs1 mutations, all confirmed to have low germination stimulant activity in agar assays. The findings demonstrate the effectiveness of integrating molecular tools into breeding programs, accelerating the development of Striga-resistant sorghum varieties and aiding genetic research into plant-parasite interactions. These advancements provide a sustainable pathway to mitigate Striga‘s devastating agricultural impact.
SorghumBase examples:
Reference:
Adeyanju AO, Rich PJ, Ejeta G. A powerful molecular marker to detect mutations at sorghum LOW GERMINATION STIMULANT 1. Plant Genome. 2024 Oct 2:e20520. PMID: 39358304. doi: 10.1002/tpg2.20520. Read more
Related Project Websites:
- Purdue Center for Global Food Security: https://www.purdue.edu/discoverypark/food/directory/executive-director.php