This study identifies novel and known genetic loci associated with sorghum root system architecture traits, highlighting their role in drought tolerance and demonstrating the potential of genomic selection for sorghum improvement.
Keywords: GWAS, Genomic prediction, Quantitative trait locus, Root system architecture, Sorghum
Scientists from Swedish University of Agricultural Sciences, Addis Ababa University, Washington State University, Bio and Emerging Technology Institute, Ethiopian Institute of Agricultural Research and Purdue University investigated the genetic basis of root system architecture (RSA) traits in sorghum, a key factor in improving crop tolerance to abiotic stresses such as drought and nutrient scarcity. Using genome-wide association studies (GWAS), novel and previously reported single nucleotide polymorphisms (SNPs) associated with RSA traits were identified. Notably, five SNP loci related to nodal root angle were mapped across chromosomes 1, 3, 6, and 7, with some co-located within genes involved in plant growth regulation and stress responses. One key SNP on chromosome 6 (sbi20340807) was linked to the gene Sobic.006G106200, which regulates chlorophyll degradation and water use efficiency, potentially contributing to the stay-green trait and improved drought tolerance. Additional SNPs on chromosomes 3 and 7 were associated with genes encoding proteins involved in root development, ethylene biosynthesis, and stress adaptation. Moreover, loci associated with root number and nodal root length were identified, including novel SNPs on chromosome 9 near genes implicated in drought response and hormonal regulation, suggesting a complex genetic control of RSA traits.
Genomic prediction models were applied to assess the heritability of RSA traits, utilizing Bayesian approaches and RR-BLUP models. Prediction accuracies ranged from 0.30 to 0.63, comparable to previous studies in other crops, with the highest accuracy observed for nodal root length. These findings highlight the potential of genomic selection in breeding programs aimed at improving sorghum’s drought resilience. However, the study acknowledges limitations, including the controlled greenhouse environment and the need for multi-environment validation. Future research should incorporate diverse genetic backgrounds and field-based phenotyping to enhance the applicability of these findings.
SorghumBase examples:
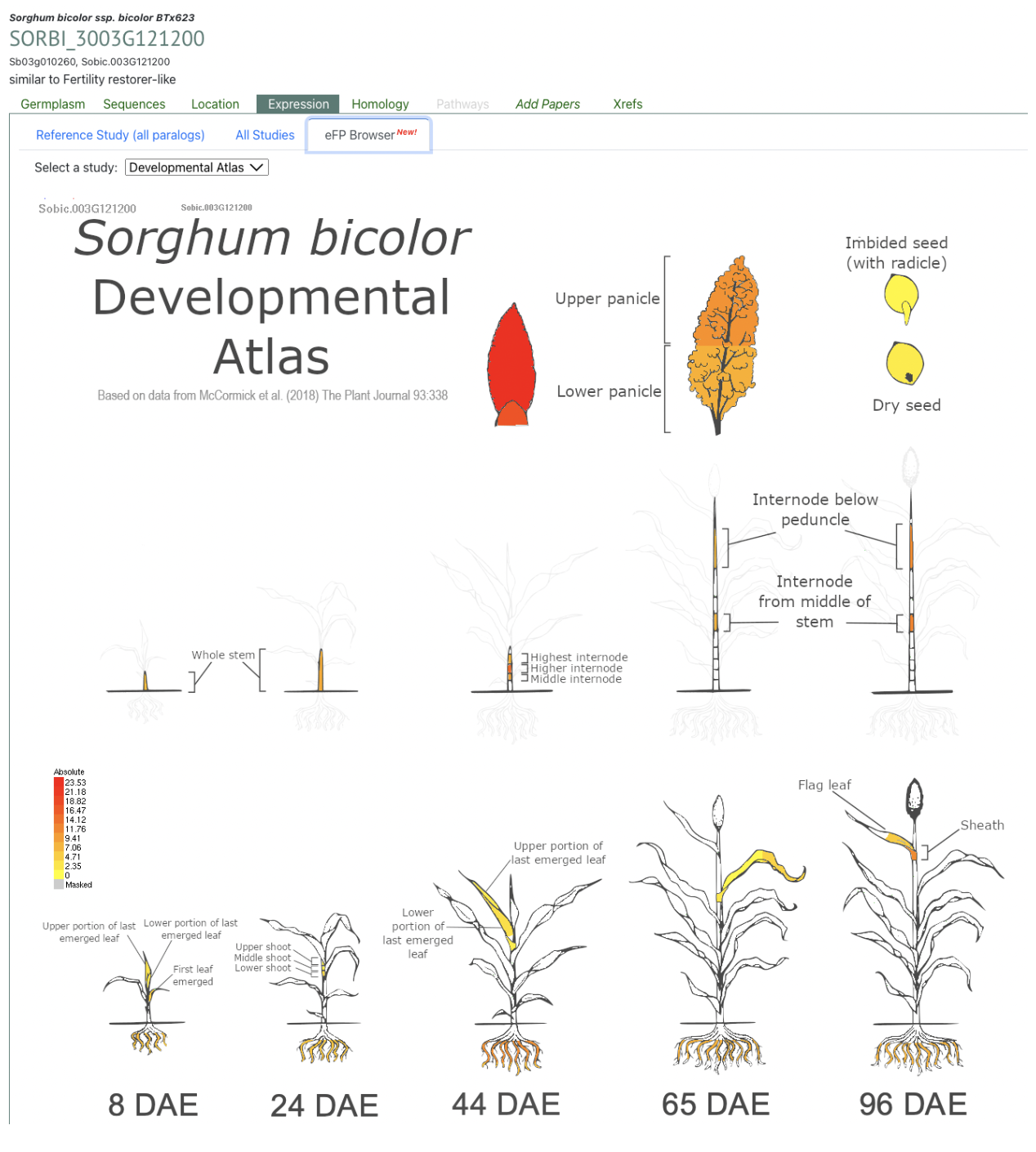
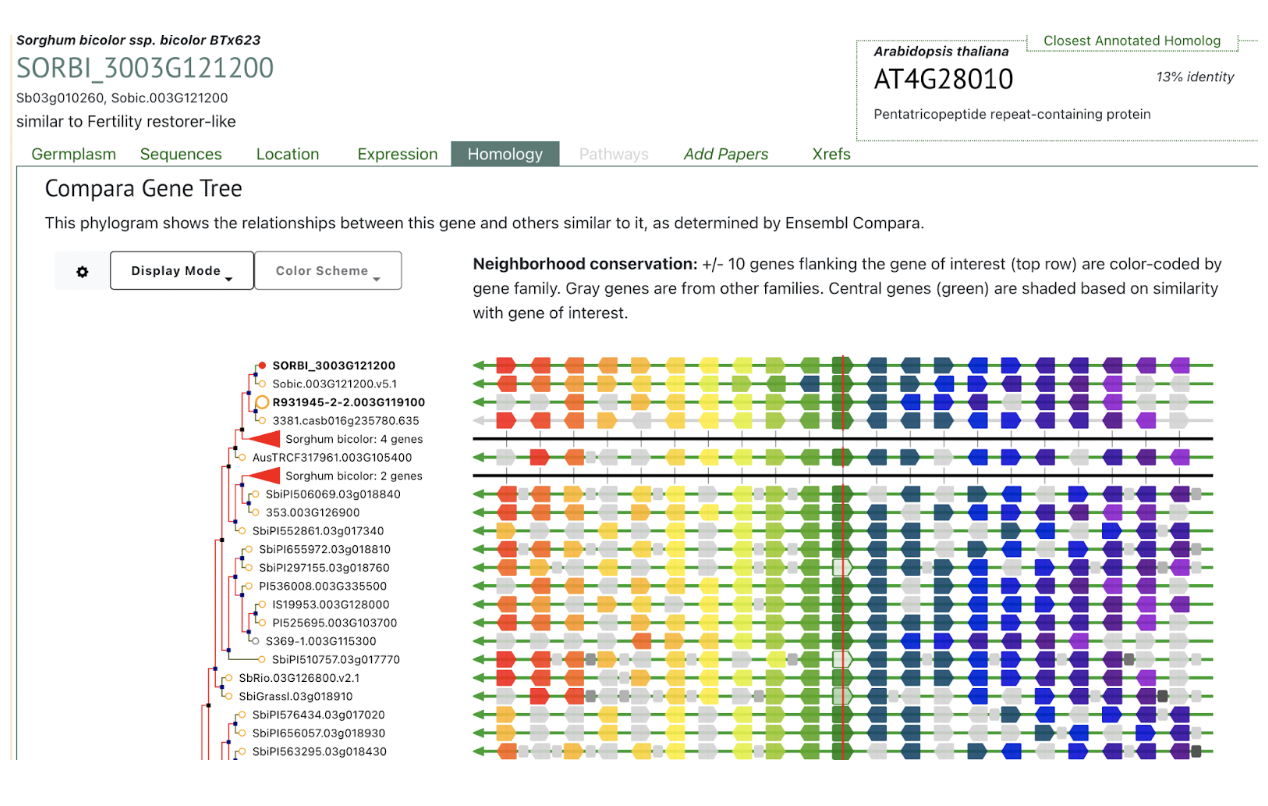

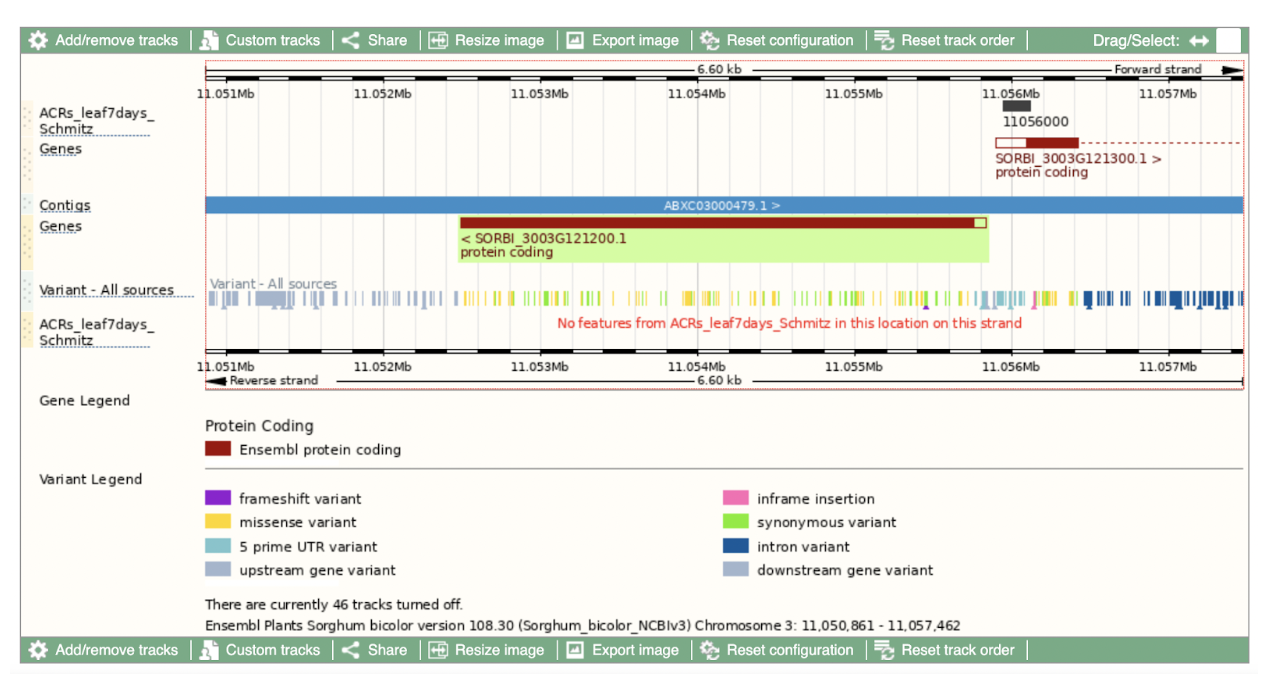
Reference:
Enyew M, Geleta M, Tesfaye K, Seyoum A, Feyissa T, Alemu A, Hammenhag C, Carlsson AS. Genome-wide association study and genomic prediction of root system architecture traits in Sorghum (Sorghum bicolor (L.) Moench) at the seedling stage. BMC Plant Biol. 2025 Jan 17;25(1):69. PMID: 39819271. doi: 10.1186/s12870-025-06077-w. Read more
Related Project Websites:
- Dr Carlsson’s page at Swedish University of Agricultural Sciences: https://www.slu.se/cv/anders-carlsson/