This study reveals that increasing m6A RNA modifications through SbMTA overexpression enhances salt tolerance in sorghum by stabilizing stress-responsive transcripts, whereas reducing m6A levels with SbALKBH10B overexpression diminishes this resilience.
Keywords: N 6-methyladenosine, SbALKBH10B, SbMTA, Epitranscriptomic regulation, salt tolerance, sorghum
Scientists from Shandong Normal University and and Xinjiang Institute of Ecology and Geography at the Chinese Academy of Sciences explored the role of m6A RNA modifications in enhancing salt tolerance in sorghum, focusing on the opposing effects of two regulatory genes, SbMTA (a writer) and SbALKBH10B (an eraser). m6A is a prevalent RNA modification in plants, playing a significant role in stress responses by altering gene expression. The researchers found that overexpressing SbMTA, which adds m6A modifications, led to increased salt tolerance in sorghum. This tolerance was associated with improved growth under salt stress, elevated m6A levels, enhanced potassium uptake, sodium exclusion, and increased stability of key stress-responsive transcripts. By contrast, overexpressing SbALKBH10B, which removes m6A modifications, decreased sorghum’s salt tolerance, indicating that m6A enrichment is beneficial for salt stress resilience.
The study highlights that SbMTA overexpression improved the stability of transcripts in pathways critical for stress responses, such as ABA signaling and ROS scavenging. In particular, transcripts like SbSNRK2.6 and SbTPPI, involved in salt and drought responses, showed increased m6A modifications, promoting their stability. These modifications supported improved ABA and auxin signaling, critical for maintaining root growth and stomatal function under stress. Conversely, decreased m6A levels in SbALKBH10B-overexpressed lines reduced these stabilizing effects, diminishing salt tolerance. These findings suggest that targeted m6A modifications mediated by SbMTA may serve as a promising genetic approach to enhance sorghum’s resilience to salt stress, with potential applications for improving crop stress tolerance more broadly.
SorghumBase example:
SbMTA (SORBI_3004G285000)
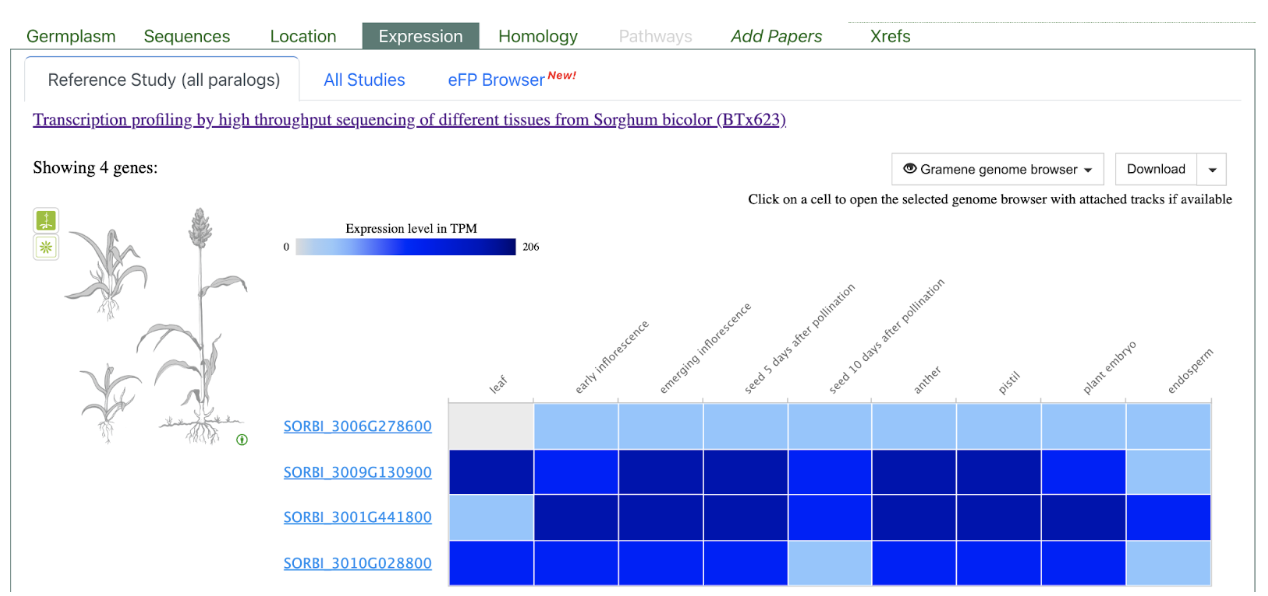
SbALKBH10B (SORBI_3006G278600)
Salt Tolerance QTL (Relative salt injury rate)
Reference:
Zheng H, Dang Y, Gao Y, Li S, Wu F, Zhang F, Wang X, Du X, Wang L, Song J, Sui N. An mRNA methylase and demethylase regulate sorghum salt tolerance by mediating N6-methyladenosine modification. Plant Physiol. 2024 Oct 15:kiae529. PMID: 39405192. doi: 10.1093/plphys/kiae529. Read more