This study identifies key transcription factors, metabolic pathways, and hormone signaling mechanisms that differentiate salt-tolerant and salt-sensitive sorghum cultivars, providing insights for breeding salt-resistant varieties.
Keywords: Brewing sorghum, Metabolome, Salinity tolerance, Transcription factor, Transcriptome
Soil salinity remains a significant threat to crop production and food security, necessitating the cultivation of salt-tolerant crop varieties. Scientists from Agricultural College of Inner Mongolia Minzu University and Tongliao Agriculture and Animal Husbandry Research Institute examined the physiological, transcriptomic, and metabolomic responses of two brewing sorghum cultivars with differing salt tolerance levels. Transcription factors (TFs) play a crucial role in regulating stress responses, with families such as MYB, NAC, and WRKY exhibiting differential expression under salt stress. MYB TFs regulate key genes involved in stress adaptation, while NAC TFs, known for their role in abiotic stress tolerance, showed cultivar-specific expression patterns. Similarly, WRKY TFs, widely implicated in plant defense and development, displayed differential expression, suggesting distinct regulatory strategies in salt-tolerant (NY) and salt-sensitive (MY) cultivars. These findings highlight the potential of manipulating TFs to enhance salt tolerance in sorghum breeding programs.
Metabolomic analysis revealed significant metabolic reprogramming in response to salinity stress. The salt-tolerant cultivar NY exhibited up-regulation of metabolites linked to osmotic adjustment and stress defense, such as phenylpropanoids, amino acids, and unsaturated fatty acids, contributing to membrane stability and oxidative stress protection. In contrast, the salt-sensitive MY cultivar showed metabolic shifts focused on central metabolism pathways but lacked efficient stress mitigation. Additionally, hormone-related gene expression varied between cultivars, with auxin-responsive genes showing differential regulation, indicating their role in salt adaptation. The study underscores the importance of transcription factors, metabolic pathways, and hormone signaling in mediating salt stress responses. These findings provide valuable molecular targets for breeding salt-tolerant sorghum varieties, offering a pathway toward improved crop resilience in saline environments.
SorghumBase examples:
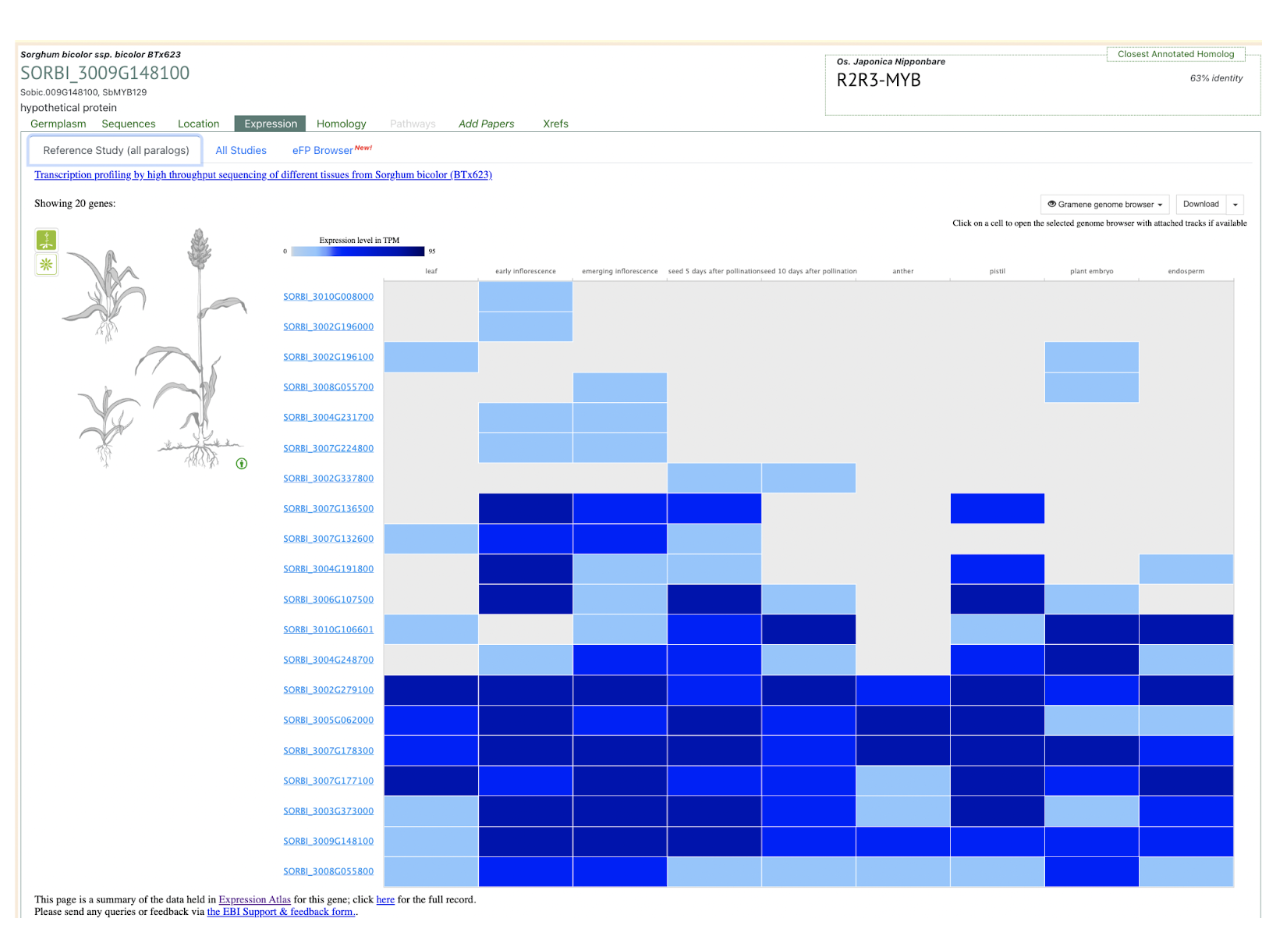
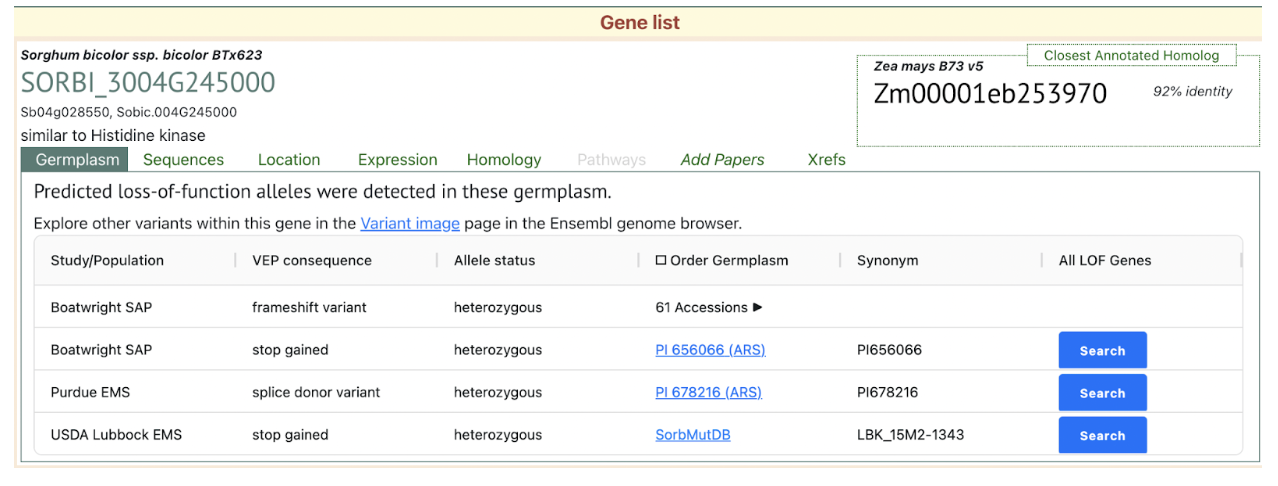
Reference:
Zhou W, Wang ZG, Li Y, Wu GJ, Li M, Deng ZL, Cui FJ, Xu QQ, Li Y, Zhou YX. Comparative transcriptome and metabolome analysis reveals the differential response to salinity stress of two genotypes brewing sorghum. Sci Rep. 2025 Jan 27;15(1):3365. PMID: 39870699. doi:10.1038/s41598-025-87100-w. Read more